Cave bears were giant plant eating bears that roamed Europe and northern Asia, and went extinct around 25 thousand years ago. They hibernated in caves during the winter. This was a dangerous time, as those which had failed to fatten up enough during the summer would not survive hibernation.
As a result, many caves across Europe and northern Asia are now filled with the bones of cave bears, each one containing potentially thousands of individuals. In our new study, we analysed a bone from a cave in the Caucasus Mountains.
Our team recovered the genome from a 360,000-year-old cave bear, revealing new details of the animals’ evolutionary history and almost rewriting their entire evolutionary tree. As well as what it can tell us about cave bear evolution, this discovery is a breakthrough for the field of ancient DNA.
February 2021 was an important month for the study of palaeogenomes – the analysis of genomes from extinct species. Two studies were published just one day apart, one reporting the oldest genome from a permafrost environment – from a 1.2 million year old mammoth tooth – and our new research, reporting the oldest genome from a non-permafrost environment.
Non-permafrost genomes

Johanna Paijmans, Author provided
After death, the environment in which an animal dies in strongly affects the speed at which its tissues degrade. We see this every day in our kitchens – food left out on a hot day quickly spoils, but the same food stored in the freezer can last for months. DNA is no different, it survives a long time in the near perfect conditions of permafrost. But the warmer the storage conditions, like in non-permafrost environments, the faster DNA will degrade to a state where it’s no longer recognisable as the original product.
Even if the DNA has survived all that time, a major challenge for palaeogenome recovery is ancient samples are also usually highly contaminated with microbial DNA from the external environment – like the bacteria that fed on the decaying corpse or live in the surrounding soil. These typically outnumber the sample DNA, which increases the cost of genome sequencing by up to a factor of 100. This make most palaeogenome sequencing of ancient samples simply too expensive to undertake.
Read more:
We sequenced the oldest ever DNA, from million-year-old mammoths
But a recent discovery in the field of ancient DNA has changed things. A particular bone in the mammalian skeleton – the petrous bone, part of the skull which contains the inner ear – seems to be more resistant to contamination from the external environment, potentially due to its extremely high density. In our previous studies we used petrous bones to sequence the genomes of much younger extinct cave bears. Some of these have contained as much a 70% cave bear DNA.
We wondered if this was the way to recover even older palaeogenomes. So we selected a cave bear petrous bone dating to 360,000 years ago, within the geological period known as the Middle Pleistocene, and took it to the lab.
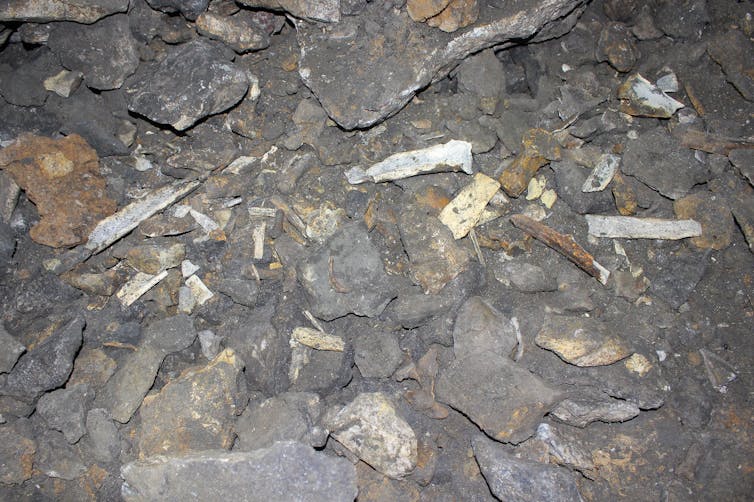

Axel Barlow., Author provided
Failures
Our first attempt in the lab was actually a complete failure. We extracted DNA from the petrous bone and produced around a million DNA sequences. Only 74 out of the million provided a match to the polar bear genome, which we use as a reference to identify and assemble the short sequences of cave bear DNA.
Undeterred, we tried again, and this time we hit the jackpot. Tens of thousands of sequences, around 3.6% of the data, showed matches to the polar bear. In total, we produced around 2.1 billion base pairs – individual pieces of genetic code – of the genome of this ancient cave bear.
Palaeontological and ancient DNA research has found distinct groups of cave bears, some even regarded as distinct species. Much of this genetic evidence comes from maternally inherited mitochondrial DNA, which represents only a tiny part of an animal’s total DNA.
We constructed an new evolutionary family tree of cave bears from their entire nuclear genomes and compared it to the same tree generated from mitochondrial DNA. To our surprise, the two trees were almost completely different. In a single analysis we had essentially rewritten our understanding of cave bear evolution. It turned out the evolutionary relationships between cave bears based on the maternal line alone are very different than when looking at their entire DNA.
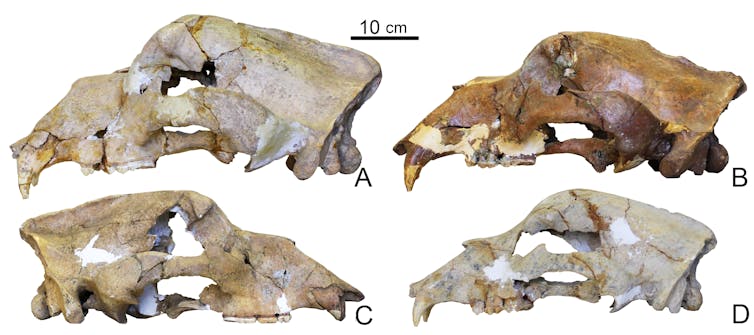

Gennady Baryshnikov, Author provided
Our results showed the three main lineages of cave bear started to diverge around one million years ago, at the same time as their nearest relatives, polar bears and brown bears, diverged from each other. Interestingly, this was a time when glacial cycles – “ice ages” – became longer and more intense, which we think may have been a factor driving the evolution of these bears.
Although only based on a single sample, our study represents a big technological achievement. It suggests petrous bones offer exciting potential to further push back the time limits of palaeogenome recovery across all environments and ecosystems.
The petrous bone is present in all mammals, so it’s of particular importance for mammalian biodiversity. For non-mammalian remains, we’ve yet to find the best element for palaeogenomic analysis – but based on our results, we know we should be looking for the densest parts of the skeleton.
The hot and humid tropical environments, which present the most challenging conditions for DNA preservation, are generally considered biodiversity hotspots – where the majority of the world’s biodiversity is found. These regions may represent the next frontier in palaeogenome research, to explore that unknown past genetic biodiversity hidden in bones.
![]()
Axel Barlow receives funding from The Natural Environment Research Council and the European Research Council (EasiGenomics).
Johanna L.A. Paijmans does not work for, consult, own shares in or receive funding from any company or organisation that would benefit from this article, and has disclosed no relevant affiliations beyond their academic appointment.